Lipid membranes and cell-free reconstitution
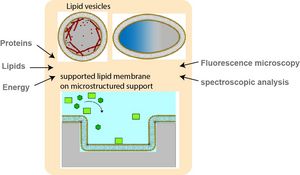
The reconstitution of cellular functions from the scratch (by using purified proteins, DNA and lipids) is an intriguing approach to study the molecular processes, that underly cellular functions, in well-defined environments. Our group builds biomimicries with high control over system parameters and reconstitutes the assembly of protein structures and patterns. We are particularly focusing on the reconstitution of cellular morphogenesis, which plays a key role in many biological and biomedical contexts, including cell migration, immunological responses, cancer metastases and tissue regeneration. Our toolbox includes various lipid membrane systems, protein purification techniques, fluorescence microscopy and analysis methods.
Selected publications:
Zieske K, Chwastek G, Schwille P (2016) Protein Patterns and Oscillations on Lipid Monolayers and in Microdroplets. Angew Chem Int Ed Engl. 55(43):13455-13459. [MPI for Biochemistry, Martinsried]
Zieske K, Schwille P. (2014) Reconstitution of self-organizing protein gradients as spatial cues in cell-free systems. Elife.3. doi: 10.7554/eLife.03949. [MPI for Biochemistry, Martinsried]
Zieske K, Schwille P. (2013) Reconstitution of pole-to-pole oscillations of min proteins in microengineered polydimethylsiloxane compartments. Angew Chem Int Ed Engl. 52(1):459-62. [MPI for Biochemistry, Martinsried]
Contact
Zieske Research Group
MPI for the Science of Light
Staudtstr. 2
D-91058 Erlangen, Germany





